Why Can Evolution in Mosquitoes Be More Easily Observed Than Evolution in a Population of Squirrels?
Abstract
Living with relatives can exist highly beneficial, enhancing reproduction and survival. High relatedness can, even so, increase susceptibility to pathogens. Here, nosotros examine whether the benefits of living with relatives offset the harm caused by pathogens, and if this depends on whether species typically live with kin. Using comparative meta-analysis of plants, animals, and a bacterium (due north species = 56), we show that loftier inside-group relatedness increases mortality when pathogens are present. In contrast, mortality decreased with relatedness when pathogens were rare, peculiarly in species that live with kin. Furthermore, beyond groups variation in mortality was lower when relatedness was loftier, simply abundances of pathogens were more than variable. The effects of inside-group relatedness were only evident when pathogens were experimentally manipulated, suggesting that the harm caused past pathogens is masked by the benefits of living with relatives in nature. These results highlight the importance of kin selection for understanding disease spread in natural populations.
eLife digest
Living in a grouping with relatives has many advantages, such every bit helping with child rearing and gathering food. This has led many species to evolve a range of grouping behaviours; for example, in dearest bee populations, worker bees sacrifice themselves to relieve the colony from incoming enemies.
Just there are also downsides to living with family. For example, bacteria, viruses and other illness-causing pathogens will find it easier to spread between relatives. This is because individuals with the same genes have similar immune defences. So, is information technology better to live with relatives who can help with life's struggles or alive with unrelated individuals where at that place is a lower chance of getting ill?
To aid answer this question, Bensch et al. analysed data from 75 studies which encompassed 56 different species of plants, animals, and one type of leaner. This showed that creatures living in family groups experienced more disease and had a higher risk of death. However, if groups had a low take a chance of encountering pathogens, individuals living with relatives were more than likely to survive. This cancels out the disadvantages family groups confront when pathogens are more than common.
The analysis by Bensch et al. provides new insights into how pathogens spread in species with different social systems. This information tin be used to predict how diseases occur in nature which will benefit ecologists, epidemiologists, and conservation biologists.
Introduction
Loftier relatedness between individuals can favour the evolution of cooperative interactions that increase reproductive success and survival (Hamilton, 1964a; Hamilton, 1964b). For example, it has been repeatedly shown that individuals can laissez passer on their genes indirectly by providing vital resources to relatives and assisting them with tasks that are difficult to practice lonely, such as caring for offspring (Alexander, 1974; Rubenstein and Abbot, 2017; West et al., 2007). However, living with relatives can also increase susceptibility to pathogens that spread more hands amongst genetically like individuals, with similar immune defences (Anderson et al., 1986; Baer and Schmid-Hempel, 1999; Hamilton, 1987; Liersch and Schmid-Hempel, 1998; Schmid-Hempel, 1998; Sherman et al., 1998). This phenomenon has been referred to equally the 'monoculture event' (Elton, 1958) in agricultural settings after it was observed that clonal crops were highly susceptible to illness outbreaks (Garrett and Mundt, 1999; Tooker and Frank, 2012; Wolfe, 1985; Zhu et al., 2000). More than recently it has as well been established that such effects occur in natural populations, with higher genetic similarity between individuals increasing rates of parasitism (Ekroth et al., 2019). What remains unclear is whether this translates into higher rates of mortality, or whether the benefits of living with relatives are large enough to offset the costs of increased affliction adventure (Hughes et al., 2002).
Previous research into the effects of relatedness on disease spread take been conducted on an expansive range of species including bacteria, plants, and animals. These studies have revealed remarkable variation in how changes in relatedness influence parasitism and bloodshed. For example, in honeybees, Apis melifera, high relatedness among individuals increases the risk of affliction and colony expiry (Tarpy et al., 2013), whereas in Pharoah ants, Monomorium pharaonis, high relatedness reduces the affluence of pathogens (Schmidt et al., 2011). Such differences betwixt species may in part exist due to how data are collected. In some studies, relatedness and pathogens have been experimentally manipulated, whereas in other studies relatedness and the abundances of pathogens are just observed ('observational studies'). In observational studies, results can exist variable and difficult to interpret because the causality behind relationships is uncertain (Lively et al., 2014). For case, a negative relationship betwixt relatedness and the abundance of pathogens can occur either because groups of relatives are less susceptible to pathogens, or because groups of relatives dice from pathogens and then are rarely observed (Ben-Ami and Heller, 2005; Male monarch et al., 2011; Instructor et al., 2009).
Additionally, species may vary in their susceptibility to pathogens because of differences in past selection to command affliction spread amid individuals (Loehle, 1995; Romano et al., 2020). In species where relatives ofttimes interact, selection is predicted to favour the evolution of strategies that mitigate the impacts of pathogens (Loehle, 1995; Romano et al., 2020). Limiting social interaction through group-level organization, such as job partitioning and other mechanisms of the so-chosen 'social amnesty', tin forbid disease spread among relatives (Camargo et al., 2007; Cremer and Sixt, 2009; Liu et al., 2019; Ugelvig et al., 2010; Waddington and Hughes, 2010). However, whether species that typically live with kin are better able to cope with pathogens when relatives interact, compared to species that alive with non-kin, is unclear.
The spread of illness through populations besides depends on how variable pathogen abundances are across groups. Pathogen abundances are expected to be more variable among groups of relatives because they either incorporate resistant or susceptible genotypes (Boomsma and Ratnieks, 1996; van Baalen and Beekman, 2006). Groups of unrelated individuals, on the other hand, volition contain a mix of susceptible and resistant genotypes, leading to more than predictable pathogen abundances and rates of mortality beyond groups. Such differences in variation beyond groups of related and unrelated individuals are nevertheless predicted to depend on pathogen multifariousness. When there are many unlike pathogens, groups of relatives are more likely to be susceptible to at least 1 pathogen, which can reduce variation in total pathogen abundance to a level that is similar to groups of unrelated individuals (Ganz and Ebert, 2010; van Baalen and Beekman, 2006). While both increases and decreases in variation in rates of parasitism and bloodshed have been found in specific report species (Ganz and Ebert, 2010; Johnson et al., 2006; Seeley and Tarpy, 2007; Thonhauser et al., 2016), whether variation among groups of relatives is mostly college beyond species remains to be tested.
Here, nosotros use phylogenetic meta-analysis to first examine whether the benefits of living with relatives counteract the costs of increased susceptibility to pathogens. 2nd, we tested if the power to observe such effects is dependent upon the experimental manipulation of pathogens and inside-grouping relatedness. Third, we examined if species that typically live with kin have evolved mechanisms to reduce pathogen spread among relatives compared to species that typically alive with non-kin. Finally, we investigated whether variation in the abundance of pathogens and rates of mortality is higher beyond groups of relatives. The influence of relatedness on mortality and pathogen abundances were quantified past extracting issue sizes (Pearson's correlation coefficients r) from 75 published studies across 56 species (Supplementary file 1—Tables S1-S3). Variation in pathogen abundances and rates of bloodshed were measured using a standardised effect size of variance that accounts for differences in ways, the coefficient of variation ratio (CVR), which was possible to gauge for 25 species (Supplementary file ane—Table S4).
Results
Relatedness and susceptibility to pathogens
Across species, inside-group relatedness had highly variable effects on rates of mortality and the abundance of pathogens (Effigy 1. Bayesian Phylogenetic Multi-level Meta-regression (BPMM): Mean effect size [posterior mode (PM)] of Zr = 0.06, credible interval [CI] = −0.12 to 0.26, pMCMC = 0.40. Supplementary file 1—Tabular array S5). Such variation was ubiquitous across all taxonomic groups and was largely independent of phylogenetic history (% of variation in Zr explained by phylogeny PM (CI) = eight.20 (0.11, 31.59). Figure i; Supplementary file 1—Table S5). Bloodshed was, withal, consistently higher in groups of relatives in the presence of pathogens compared to when they were absent (Figure 2. Zr pathogens absent versus nowadays PM (CI) = −0.29 (−0.44, –0.12), pMCMC = 0.002. Supplementary file one—Table S6). Similar furnishings of within-group relatedness on pathogen abundances were found (Zr pathogen affluence versus mortality PM (CI) = 0.01 (–0.ten, 0.19), pMCMC = 0.51. Supplementary file one—Table S6), but these were much weaker (Zr pathogen abundance PM (CI) = 0.ten (–0.10, 0.33), pMCMC = 0.31. Supplementary file one—Table S6).
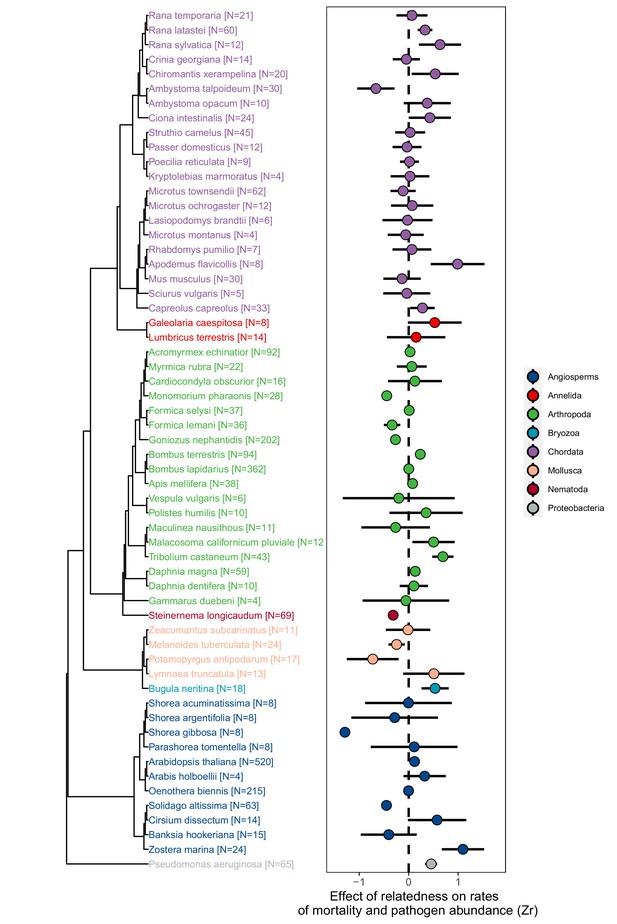
The consequence of relatedness on rates of mortality and pathogen affluence across animals, plants, and leaner.
Positive consequence sizes (Zr) indicate that mortality and/or pathogen abundances increase with the levels of relatedness within groups, negative values show decreases, and values of cipher (dotted line) are where there was no relationship. Points stand for weighted means for each species and bars are 95% confidence intervals calculated from the sample sizes of the number of groups studied which are given in brackets. See Figure 1—figure supplements 1–3 for data on obtaining event size information and testing for publication bias.
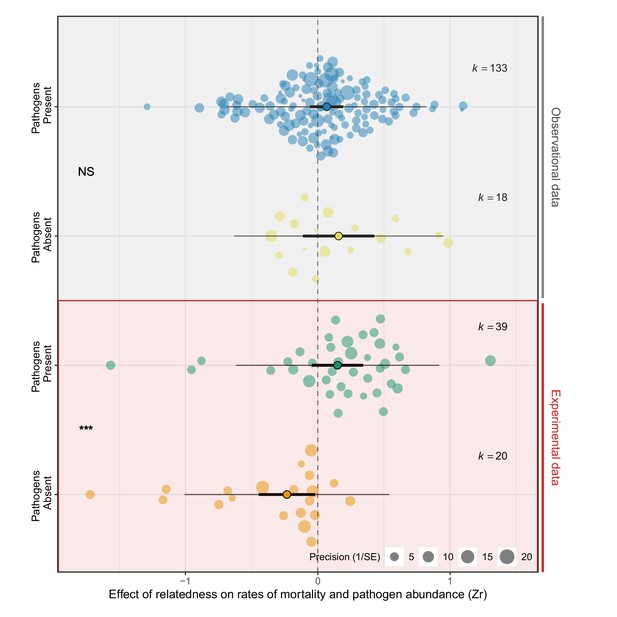
Experimental manipulations are cardinal to detecting the effects of pathogens on groups of relatives.
Positive effect sizes (Zr) indicate that mortality and/or pathogen abundances increase with the levels of relatedness within groups, negative values show decreases, and values of aught (dotted line) are where there was no human relationship. Studies that experimentally manipulated pathogen presence showed that groups of relatives had higher rates of mortality when pathogens were present, only lower bloodshed when pathogens were absent. Points with black edges correspond means, thick bars are 95% CIs, thin bars are prediction intervals, and grand is the number of effect sizes. Each dot is an private effect size and with size scaled to 1/SE (orchard plots: Nakagawa et al., 2021). Statistical differences are from Bayesian Phylogenetic Multi-level Meta-regressions (BPMMs) and placed mid-mode betwixt comparison groups denoted with symbols: NS = non-significant, *pMCMC < 0.05, **pMCMC < 0.01, ***pMCMC < 0.001.
Experimental studies reveal contrasting effects of relatedness in the presence and absenteeism of pathogens
There was evidence that pathogens causally increased bloodshed in groups of relatives (Figure 2; Supplementary file 1—Table S7). In studies where pathogens were experimentally manipulated, groups of relatives had significantly college bloodshed when pathogens were present compared to when they were absent (Zr pathogens absent versus present PM (CI) = −0.40 (−0.57, –0.21), pMCMC = 0.001. Effigy 2; Supplementary file 1—Table S7). The contrasting effects of relatedness in the presence and absence of pathogens meant that overall the result of relatedness on mortality did not significantly differ from zero (Zr pathogens nowadays PM (CI) = 0.17 (−0.09, 0.38), pMCMC = 0.fifteen. Zr pathogens absent PM (CI) = −0.23 (−0.49, 0.03), pMCMC = 0.eleven. Figure 2; Supplementary file 1—Table S7). Therefore the greater susceptibility of groups of relatives to pathogens appears to be masked by kin selected benefits of living with relatives when pathogens are rare. This may also explain why in observational studies the consequence of relatedness on mortality, both in the presence and absenteeism of pathogens, was shut to zero (Zr pathogens present PM (CI) = 0.06 (−0.10, 0.27), pMCMC = 0.36. Zr pathogens absent-minded PM (CI) = 0.10 (−0.xi, 0.50), pMCMC = 0.26. Figure 2; Supplementary file 1—Tabular array S7).
Experimental manipulations of relatedness were less important for detecting the effects of relatedness on bloodshed than manipulations of pathogens (Supplementary file 1—Table S8). Studies that experimentally manipulated inside-grouping relatedness found similar reductions in survival in groups of relatives when pathogens were present to observational studies (Experimental studies: Zr pathogens absent versus present PM (CI) = −0.19 (−0.42, –0.08), pMCMC = 0.01. Observational studies: Zr pathogens absent-minded versus nowadays PM (CI) = −0.xxx (−0.77, –0.10), pMCMC = 0.02. Supplementary file 1—Table S8).
Responses to pathogens depend on whether species alive in kin groups
Next, nosotros tested whether species that typically live with relatives have evolved mechanisms to limit the negative effects of pathogens when inside-group relatedness is high. To do this, species that typically live with relatives under natural conditions (r > 0.25 referred to every bit 'kin') were compared to those that associate with unrelated individuals (r < 0.25 referred to as 'non-kin'. See Materials and methods for details of data used to allocate species). When pathogens were present, the effect of relatedness on rates of mortality did not differ between species that alive with kin and non-kin (Zr kin versus not-kin pathogen present PM (CI) = 0.09 (−0.31, 0.37), pMCMC = 0.79. Supplementary file 1—Table S9). However, when pathogens were absent, high relatedness reduced mortality in species that alive with kin, but increased mortality in species that alive with non-kin (Zr kin versus not-kin pathogen absent PM (CI) = −0.57 (−ane.eleven, 0.02), pMCMC = 0.03. Effigy 3, Supplementary file 1—Table S9). For example, in the ruby-red flour beetle, Tribolium castaneum, and the tube worm, Galeolaria caespitosa, that typically collaborate with non-kin, mortality was ii to iv times higher when individuals were placed in groups of relatives compared to when individuals were unrelated (Agashe, 2009; McLeod and Marshall, 2009). These results prove that species that typically associate with not-kin endure reductions in fitness when placed in groups of relatives, merely only when pathogens are rare. Conversely, species that live with kin have higher fettle in groups of relatives when pathogens are absent, but such benefits disappear when pathogens are present (Zr pathogens absent versus present PM (CI) = −0.33 (−0.53,–0.16), pMCMC = 0.001. Supplementary file 1—Table S9).
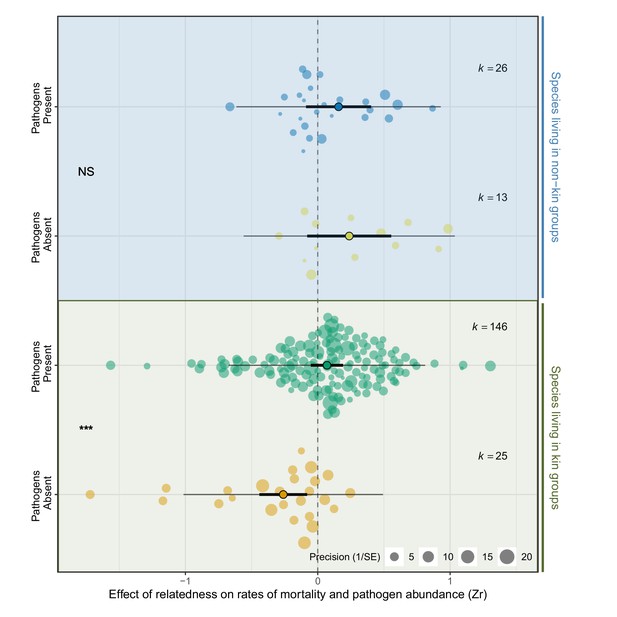
Species that live in kin groups responded differently to experimental manipulation of pathogens compared to species that live with non-kin.
Positive outcome sizes (Zr) indicate that mortality and/or pathogen abundances increase with the levels of relatedness within groups, negative values show decreases, and values of zippo (dotted line) are where there was no relationship. When pathogens were experimentally removed species that live with kin had higher survival, which was reversed when pathogens were present. In contrast, in that location was no effect of relatedness on bloodshed when pathogens were present or absent-minded in species that live with not-kin. The components of the orchard plots are the same as in Figure ii. Statistical differences are from Bayesian Phylogenetic Multi-level Meta-regressions (BPMMs) and placed mid-way betwixt comparing groups denoted with symbols: NS = not-significant, *pMCMC < 0.05, **pMCMC < 0.01, ***pMCMC < 0.001.
Relatedness increases variance in mortality beyond groups, simply not pathogen abundances
Variation in rates of bloodshed and the abundance of pathogens were influenced by relatedness in opposing ways (Figure 4; Supplementary file one—Table S10-S13). Experimental manipulations of pathogen presence were of import for detecting these effects (Supplementary file 1—Tabular array S12). In observational studies, relatedness inside groups had no effect on variance in bloodshed, either in the presence or absence of pathogens, and did not influence variance in pathogen abundances (Mortality pathogens absent: LnCVR PM (CI) = 0.44 (−0.32, 0.94), pMCMC = 0.32. Mortality pathogens present: LnCVR PM (CI) = −0.03 (−0.74, 0.65), pMCMC = 0.93. Pathogen abundance: LnCVR PM (CI) = −0.15 (−0.71, 0.fifty), pMCMC = 0.83. Effigy 4, Supplementary file 1—Tabular array S12). In contrast, in experimental studies bloodshed was more variable across groups of relatives when pathogens were present (LnCVR PM (CI) = 0.88 (0.21, 1.41), pMCMC = 0.02. Figure four, Supplementary file 1—Table S12). The opposite design was true for pathogen abundances, with groups of relatives being less variable. This meant that overall, mortality was significantly more variable than the abundance of pathogens among groups of related versus unrelated individuals (LnCVR PM (CI) = 1.18 (0.56, 1.88), pMCMC = 0.001. Figure iv, Supplementary file 1—Table S12). These results propose that pathogens spread more uniformly across groups of relatives, but effects on mortality are more variable than across groups of unrelated individuals.
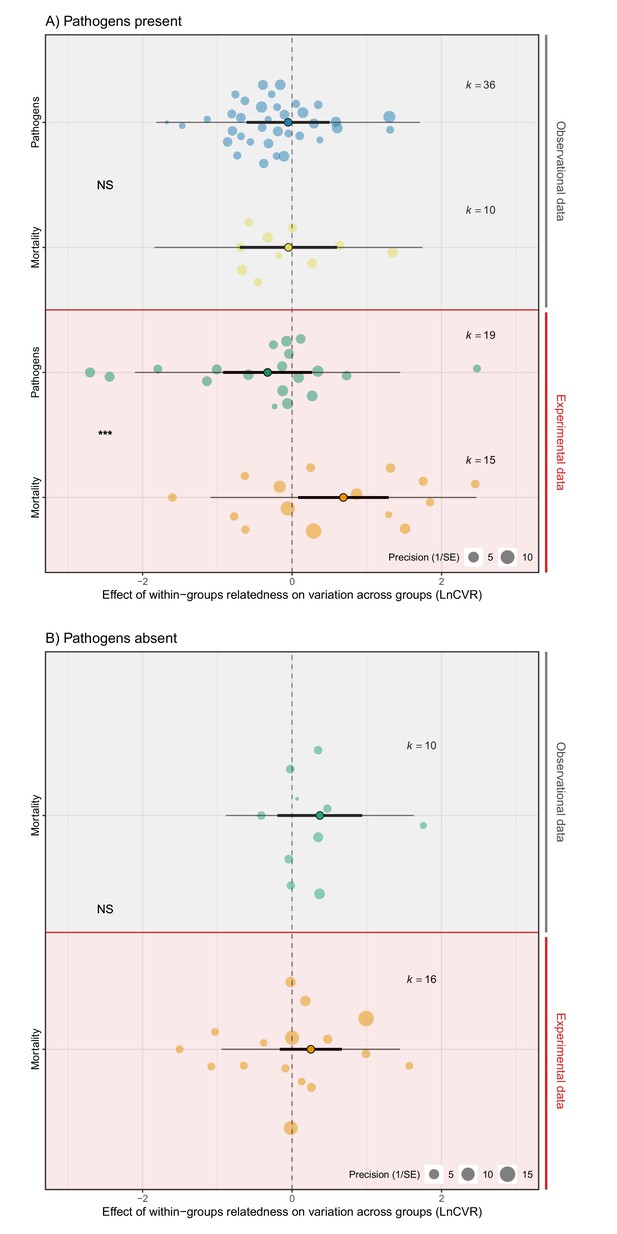
The effect of within-group relatedness on variance in mortality and pathogen abundance.
Positive effect sizes (LnCVR) show that variation in rates of bloodshed and/or pathogen abundances across groups (accounting for mean differences – see Figure 4—figure supplement i for mean variance relationship) increases with within-grouping relatedness, negative values show decreases in variance, and aught values show no change in variance (dotted line). (A) In the presence of pathogens, relatedness increased variance in mortality, just decreased variance in pathogen abundance. (B) When pathogens were absent-minded, relatedness did not influence variance in mortality. The components of the orchard plots are the same as in Effigy ii. Statistical differences are from Bayesian Phylogenetic Multi-level Meta-regressions (BPMMs) and placed mid-manner between comparison groups denoted with symbols: NS = non-significant, *pMCMC < 0.05, ** pMCMC < 0.01, ***pMCMC < 0.001. Figure iv—effigy supplement 2 for examination of publication bias.
Discussion
Our analyses show that pathogens can increment rates of mortality in groups of relatives. The detrimental effects of pathogens were, still, counteracted past high relatedness reducing bloodshed when pathogens were rare, especially in species that alive in kin groups. Such contrasting furnishings of relatedness meant that experimental manipulations were crucial for detecting the costs and benefits of living with relatives when the presence of pathogens varied. Additionally, high relatedness resulted in more even abundances of pathogens across groups, but more variable rates of mortality, highlighting the importance of population genetic structure in explaining the epidemiology of diseases. Nosotros discuss these findings in relation to the environments favouring the evolution of different social systems, the mechanisms that take evolved to forbid disease spread in social groups, and the types of study arrangement where more experimental data are required.
The interaction betwixt kin selected benefits and mortality caused by pathogens has important implications for our understanding of the ecological distributions of species and the evolutionary origins of different social systems. In some lineages, such as birds, cooperative species that live in families have been constitute to inhabit areas that are hot and dry (Cornwallis et al., 2017; Jetz and Rubenstein, 2011; Lukas and Clutton-Brock, 2017). This has been attributed to the benefits of cooperative offspring intendance existence higher in environments that are challenging for independent reproduction (Emlen, 1982). An additional, potentially of import explanation is that the costs imposed by pathogens when living with relatives may exist lower in such environments (Campbell-Lendrum et al., 2015). Parallel arguments have been made for social insects. Species with sterile worker castes, that only evolved in groups with loftier levels of relatedness, are idea to accept arisen in environments protected from pathogens (Hamilton, 1987). For example, sterile soldier castes have evolved at to the lowest degree six contained times in clonal groups of aphids, and the bulk of these cases grade galls that provide protection confronting pathogens (Hamilton, 1987; Stern and Foster, 1996). Escape from pathogens may therefore be a general feature governing the evolutionary origin, as well equally the current ecological niches, of species living in highly related groups.
The benefits of living with relatives are predicted to generate selection for increased resistance or tolerance to illness spread (Loehle, 1995; Romano et al., 2020). Adaptations to limit pathogen manual in kin groups have been documented in some species. For example, in leaf cutter ants, Acromyrmex spp., workers exterior the colony, where pathogens are more prevalent, do not enter the inner colony (Camargo et al., 2007). Contamination of nutrient by pathogens is also limited by workers exterior the colony performing dedicated tasks, such every bit foraging versus waste management (Waddington and Hughes, 2010). Irresolute the organisational structure of groups or living in smaller groups can therefore increment social distancing and reduce pathogen transmission (Loehle, 1995; Romano et al., 2020; Liu et al., 2019).
While examples of social immunity exist, at that place was piddling evidence that species that live with kin have generally evolved mechanisms to limit the harm caused by pathogens. Species that live in kin groups suffered similar reductions in survival from pathogens to species that alive with non-kin (Figure 3). Ane explanation is that individuals respond to greater pathogen pressure by forming more genetically various groups (Schmid-Hempel, 1998; Sherman and Morton, 1988). For example, increases in mating promiscuity under higher disease run a risk tin lower relatedness among offspring recruited to groups (Busch et al., 2004; Singh et al., 2015; Soper et al., 2014). Such responses tin reduce affliction spread, but also weakens option for adaptations that limit pathogen spread amongst related individuals. The relative costs of decreasing the effects of pathogens past reducing relatedness versus other mechanisms remains unclear, only may provide insight into why dissimilar responses to pathogens have evolved across species.
High relatedness was associated with higher and more than variable rates of bloodshed in the presence of pathogens, but had little effect on variation in pathogen load. Such differences may arise because pathogen abundances are oft weakly related to the virulence of pathogens (Leggett et al., 2012). Genotypes can also be equally susceptible to pathogens, but vary in their ability to clear infections, which may explain why inside-group relatedness influenced bloodshed rates without strongly affecting variation in pathogen abundances (Best et al., 2008; Howick and Lazzaro, 2014; Koskela et al., 2002).
The effects of relatedness on bloodshed rates were only evident in experiments. At that place are a number of possible, non-mutually exclusive, explanations for this. Information technology is possible that observational studies fail to capture the truthful consequence of pathogens because of sampling biases: groups of relatives infected with pathogens tin can quickly die resulting in their effects being underestimated (Ben-Ami and Heller, 2005; King et al., 2011; Instructor et al., 2009). The diversity and affluence of pathogens may also differ between experimental and observational studies. Although experiments often reported that pathogens were manipulated in biologically realistic ways, it is possible that pathogen abundances are by and large college in experiments leading to larger outcome sizes. Additionally, experiments generally only manipulated single pathogens whereas observational studies on natural populations often involve communities of pathogens. Low pathogen diversity is predicted to increase variation across groups of relatives (Boomsma and Ratnieks, 1996; van Baalen and Beekman, 2006). The lack of an effect of relatedness on variance in mortality in observational studies may therefore be due to the diversity of pathogens being college. In our dataset, there was only ane experimental written report that manipulated multiple pathogens. In Daphnia magna it was plant that variance in parasitism was higher in groups of relatives ('clonal' versus 'polyclonal' populations), but this diminished as the number of pathogens increased (Ganz and Ebert, 2010). This suggests that where pathogen variety is high, groups of relatives get increasingly susceptible to pathogens, reducing variance beyond groups (Boomsma and Ratnieks, 1996; Parsche and Lattorff, 2018; van Baalen and Beekman, 2006).
How relatedness amongst individuals influences pathogen spread has been investigated in a diverse range of species making our analyses possible. Nevertheless, experiments manipulating pathogen presence, abundance, and multifariousness across species with different ecological niches and social systems, especially those that typically acquaintance with non-kin, remain limited. In-depth analyses comparing species with ancestrally versus derived levels of high and low relatedness will also aid shed light on the importance of current versus past evolutionary responses to pathogens. We hope that our results stimulate further research in these areas which appears crucial to understanding the impact of pathogens on natural populations.
Materials and methods
Literature searches
Request a detailed protocolA systematic literature review was performed to place studies that take examined the relationship between within-grouping relatedness and rates of bloodshed or the abundance of pathogens. Ane challenge with locating relevant literature was that some studies utilize the term relatedness while others use the term genetic diversity. Genetic diversity encompasses studies that have examined within-private genetic diversity (e.g. heterozygosity), equally well as genetic diverseness of groups. The aims of our study only relate to variation in genetic diverseness of groups (relatedness). All studies where estimates of within-grouping genetic diversity were potentially influenced by within-individual genetic diversity were excluded (see below).
The literature search was performed using the Spider web of Science (WoS) including manufactures published up to the 27 July 2020. Searches were restricted to articles in English and the WoS categories were restricted to Behavorial Sciences, Environmental, Biology, Evolutionary Biological science, Ecology, Multidisciplinary Sciences, Genetics & Heredity, Biodiversity & Conservation, Entomology, Zoology as a preliminary study (Bensch MSc thesis) showed these categories to be the ones of interest. WoS searches included the following combinations of terms in the topic field: ((('genetic diversity' OR 'genetic variability' OR 'genetic diversities') AND parasite∗) OR (('genetic diversity' OR 'genetic variability' OR 'genetic diversities') AND disease∗) OR (('genetic diverseness' OR 'genetic variability' OR 'genetic diversities') AND pathogen∗) OR (('genetic diversity' OR 'genetic variability' OR 'genetic diversities') AND survival) OR (('genetic diversity' OR 'genetic variability' OR 'genetic diversities') AND mortality) OR (relatedness AND pathogen∗) OR (relatedness AND illness∗) OR (relatedness AND parasite∗) OR (relatedness AND bloodshed) OR (relatedness AND survival∗) OR 'monoculture issue' OR 'Monoculture consequence') AND (population∗ OR grouping∗ OR colony). Initial exploration of search terms included other words ('clone', 'clonal', 'social'). However, these terms inflated the number of search hits and papers with relevant data were retrieved using other terms included in our search criteria ('group', 'colony', or 'relatedness'). The search yielded a total of 4616 returns, 4615 after removing a duplicate.
To aid finding relevant papers, abstracts were downloaded and imported into R for text analysis using the quanteda package (Benoit et al., 2018). The frequency of words in each abstract was calculated and used to create a relevance score according to the number of words with positive and negative interest for this written report. The post-obit words had positive associations (listed in guild of priority): 'genetic', 'diversity', 'diversities', 'variation', 'relatedness', 'related', 'unrelatedness', 'unrelated', 'various', 'parasite', 'ectoparasite', 'ectoparasites', 'parasites', 'pathogen', 'pathogenic', 'pathogens', 'disease', 'diseases', 'diseased', 'mortality', 'survival', 'resistance', 'infection', 'infections', 'prevalence', 'tolerance', 'transmission', 'population', 'group', 'colony', 'groups', 'colonies', 'populations'. The following words had negative associations: 'human', 'humans', 'hospital', 'cancer', 'hiv', 'patients'. Papers were sorted according to their relevance scores and then manually screened to examine whether they contained data that could be used to calculate an event size of relatedness and mortality and/or pathogen abundance. We did not include studies examining the relationship between inside-group relatedness and other fitness-related measures, such as fecundity or competitive power, considering such measures are influenced past many factors other than pathogens.
We stopped screening subsequently 2102 papers as number of new papers selected for in-depth screening decreased to less than 1% per 100 references (Figure ane—figure supplement 1). In addition to WoS searches, reference lists of central studies and the papers from which we extracted event sizes were screened for relevant chief literature. PDF files of articles selected based on abstract screening were downloaded for in-depth exam of total texts. A preferred reporting items for meta-analyses diagram (Moher et al., 2009) of the literature screening process is shown in Figure 1—figure supplement two. In total our dataset consisted of 210 effect sizes from 75 studies and 56 species (Abdi et al., 2020; Agashe, 2009; Aguirre and Marshall, 2012a; Aguirre and Marshall, 2012b; Altermatt and Ebert, 2008; Anton et al., 2007; Baer and Schmid-Hempel, 2001; Baer and Schmid-Hempel, 1999; Ben-Ami and Heller, 2005; Bensch and Cornwallis, 2017; Bichet et al., 2015; Byrne and Robert, 2000; Byrne and Whiting, 2011; Melt‐Patton et al., 2017; Cook-Patton et al., 2011; Crutsinger et al., 2006; Crutsinger et al., 2008; Dagan et al., 2017; Dagan et al., 2013; de Morais, 2020; Desai and Currie, 2015; de Vere et al., 2009; Dobelmann et al., 2017; Ellison et al., 2011; Field et al., 2007; Franklin et al., 2012; Fraser et al., 2010; Gamfeldt and Källström, 2007; Ganz and Ebert, 2010; Gardner et al., 2007; He and Lamont, 2010; Hoggard et al., 2013; Hughes and Stachowicz, 2004; Hughes and Boomsma, 2006; Hughes and Boomsma, 2004; Johansson et al., 2007; Johnson et al., 2006; Kapranas et al., 2016; Keeney et al., 2009; King et al., 2011; Kotowska et al., 2010; Lambin and Krebs, 1993; Liersch and Schmid-Hempel, 1998; Mattila et al., 2012; McLeod and Marshall, 2009; Mott et al., 2019; Neumann and Moritz, 2000; Page et al., 1995; Parker et al., 2010; Parsche and Lattorff, 2018; Pearman and Garner, 2005; Reber et al., 2008; Robinson et al., 2013; Schmidt et al., 2011; Seeley and Tarpy, 2007; Sera and Gaines, 1994; Shykoff and Schmid-Hempel, 1991; Siemens and Roy, 2005; Solazzo et al., 2014; Strauss et al., 2017; Tarpy, 2003; Tarpy and Seeley, 2006; Tarpy et al., 2013; Instructor et al., 2009; Thonhauser et al., 2016; Trouvae et al., 2003; Ugelvig et al., 2010; van Houte et al., 2016; Vanpé et al., 2009; Walls and Blaustein, 1994; Wauters et al., 1994b; Weyrauch and Grubb, 2006; Winternitz et al., 2014; Woyciechowski and Król, 2001).
Overview of study blueprint and inclusion criteria
Request a detailed protocolStudies were included if they presented information on the abundance/presence of pathogens and relatedness for 4 or more groups. Relatedness was estimated from breeding designs, pedigrees, and using molecular markers. A group was defined as three or more individuals every bit it has been shown to be sufficient for group-level defences (Hughes and Stachowicz, 2004). That said, just three studies used groups with three individuals (4%) with over 93% of studies using groups with five or more than individuals. Some studies manipulated levels of relatedness by experimentally creating groups (referred to as 'experimental relatedness'), whereas other studies measured relatedness on already established groups (referred to as 'observational relatedness'). The presence and abundance of pathogens was also experimentally manipulated in some studies (referred to as 'experimental pathogens') whereas in others pathogens were measured without whatever manipulations (referred to as 'observational pathogens').
Studies on plants were included that examined the result of pathogens and herbivores, every bit information technology has previously been argued that herbivory is equivalent to parasitism (see Price, 1980; Siemens and Roy, 2005 for discussion of herbivores as pathogens). 1 study was included from unpublished information collected by the authors on ostriches, Struthio camelus (Supplementary file ane—Tables S19). Studies were excluded if they were on domestic species or where there was the potential for within-individual genetic diversity, including inbreeding, to influence estimates of within-group relatedness. In some studies, inbreeding was non explicit but potentially possible (Supplementary file 1—Tabular array S1). We tested the sensitivity of our results to any potential inbreeding furnishings past removing these effect sizes and repeating our analyses (see verification analyses; Supplementary file 1—Tables S14 and S15). If information of interest were missing in the text or figures, authors were contacted for supplementary information or description. If authors did not answer within 3 months, the effect sizes were excluded. If studies provided multiple measures of pathogen load and/or mortality, split effect sizes were extracted. Where studies presented abundances of specific pathogens as well equally total affluence of pathogens, the full was used.
Calculating the effect size of the relationship between relatedness and rates of mortality and pathogen abundances
Request a detailed protocolThe human relationship between inside-group relatedness and bloodshed and/or pathogen abundance was analysed by comparing groups with loftier and depression relatedness (relatedness every bit a categorical variable), or by analysing variation in average within-group relatedness as a continuous variable. Information from both types of study was used to calculate a standardised effect size of the correlation between within-group relatedness and bloodshed/pathogen abundance: Pearson's correlation coefficient, r. The statistical tests presented in studies were converted to r using the online meta-analysis computer (Morris, 2019) and the R package 'esc' (Lüdecke, 2019). Measures of r were transformed to Zr using 'escalc' function in the R packet metafor (Viechtbauer, 2010).
In some studies, it was not possible to obtain effect sizes direct from the statistics reported in studies, but r could be calculated from data presented in the text and/or figures in two means. First, in studies where groups with high and low relatedness were compared, means ± SD of mortality or pathogen abundances were used to calculate r. Second, in studies where descriptive statistics (eastward.grand. ways ± SD) were reported for multiple groups that varied in relatedness, nosotros conducted our own Pearson's correlations in R (see R script 'EffectSizeCalculations' and Supplementary file one—Table S2 cavalcade 'Effect size Rscript reference'). In such cases, variation in measures of relatedness, mortality, and pathogen abundances were included by creating distributions from descriptive statistics that were sampled to create 1000 datasets. For each of these 1000 datasets, r was calculated and an average taken across the g datasets.
Computing the effect size of variance in mortality and pathogen abundances across groups of related and unrelated individuals
Asking a detailed protocolThe effect of relatedness on variance in bloodshed and pathogen abundances was calculated using the natural logarithm of the ratio betwixt the coefficient of variation from groups with loftier and low relatedness (LnCVR: Nakagawa et al., 2015). LnCVR provides a standardised measure of differences in the variability of two groups accounting for differences in the ways between groups. LnCVR was used because estimates of variation increased with the mean (Effigy 4—figure supplement 1). LnCVR was calculated from studies that presented ways and SDs (converted to SD if studies presented SEs or CIs) across groups when relatedness was depression and high. This provides a standardised measure of the effect of relatedness on variability across groups, not inside groups (SDs were from across groups, not individuals).
Information on study characteristics
Request a detailed protocolFor each effect size extracted, nosotros collected data on: (1) whether pathogens were present or absent; (two) whether pathogens were experimentally manipulated; (3) whether relatedness was experimentally manipulated; (four) the method used for measuring relatedness (pedigree or molecular markers); and (5) whether pathogen abundance or mortality were measured (where survival estimates were presented, the sign of the effect size was reversed). If at that place was no mention of pathogens in the paper, then pathogens were causeless to be nowadays when studies were conducted in the field and absent if conducted in the laboratory.
Data on species characteristics
Asking a detailed protocolFor all species in our dataset nosotros searched for whether they typically associate with kin ('kin') or not ('not-kin') during the life phase that upshot sizes were measured. Species were categorised as kin if they lived in groups where r was estimated to be equivalent to 0.25 or higher and 'non-kin' if they live in groups where relatedness was estimated to be lower than 0.25 (Supplementary file 1—Table S4). Three sources of information were used to estimate levels of relatedness among individuals: (1) estimates of relatedness caused either directly from molecular genetic analyses or records of groups of individuals with known relatedness; (ii) data on the mating organization; and (3) typical dispersal patterns, equally low dispersal from groups increases relatedness. The relevant information was collected using Google Scholar including each species proper name combined with 'genetic diversity', 'relatedness', and 'grouping' every bit search terms to collect measures of within-group relatedness; 'mating system' and 'paternity' for information on mating system; and 'dispersal' and 'philopatry' for data on dispersal. The categorisation of each species as kin or non-kin forth with show and the list of literature to back up these classifications tin can exist found in Supplementary file i—Table S4 (Abdi et al., 2020; Aguirre and Marshall, 2012a; Aguirre and Marshall, 2012b; Amiri et al., 2017; Anton et al., 2007; Arnaud, 1999; Avise and Tatarenkov, 2015; Barrett et al., 2005; Bee, 2007; Beermann et al., 2015; Ben-Ami and Heller, 2005; Bryja et al., 2008; Byrne and Robert, 2000; Byrne and Whiting, 2011; Chapuisat et al., 2004; Croshaw et al., 2009; G. Crutsinger et al., 2008; Dagan et al., 2013; Dean et al., 2006; de Morais, 2020; de Vere, 2007; de Vere et al., 2009; Dobelmann et al., 2017; Edenbrow and Croft, 2012; Farentinos, 1972; Ficetola et al., 2010; Field et al., 2007; Franklin et al., 2012; Fredensborg et al., 2005; Gamfeldt and Källström, 2007; Gardner et al., 2007; Getz et al., 1993; Goldberg et al., 2013; Goulson et al., 2002; Goymann, 2009; Graham, 1941; Griffin, 2012; Haag et al., 2002; He et al., 2004; Head and Yu, 2004; Heppleston, 1972; Heske and Ostfeld, 1990; Hoffmann et al., 2003; Hoggard et al., 2009; Hughes and Stachowicz, 2004; Johnson, 2007; Johnson et al., 2006; Kapranas et al., 2016; Kawamura et al., 1991; Keeney et al., 2009; Kelly et al., 1999; Keough, 1989; Keough and Chernoff, 1987; Kimwele and Graves, 2003; Rex et al., 2011; Kozakiewicz et al., 2009; König, 1993; Lambin and Krebs, 1991; Laurila and Seppa, 1998; Lepais et al., 2010; Liker et al., 2009; Liu et al., 2013; Mackiewicz et al., 2006; McLeod and Marshall, 2009; Meling-lópez and Ibarra-Obando, 1999; Myers et al., 2011; Oettler and Schrempf, 2016; Osváth-Ferencz et al., 2017; Pai and Bernasconi, 2007; Pietrzak et al., 2010; Platt et al., 2010; Reusch et al., 1999; Rice et al., 2009; Rock et al., 2007; T. Russell et al., 2004; Schmid-Hempel and Crozier, 1999; Schmid-Hempel and Schmid-Hempel, 2000; Schmidt et al., 2011; Schmidt et al., 2016; Schradin et al., 2010; Schrempf et al., 2006; Seppä and Walin, 1996; Seppä et al., 2009; Shapiro and Dewsbury, 1986; Siemens and Roy, 2005; Simeonovska-Nikolova, 2007; Solomon et al., 2004; Stürup et al., 2014; Sutcliffe, 2010; Svane and Havenhand, 1993; Tarpy, 2003; Tatarenkov et al., 2007; Thonhauser et al., 2016; Trouvae et al., 2003; Vanpé et al., 2009; Verrell and Krenz, 1998; Walck et al., 2001; Waldman, 1982; Walls and Blaustein, 1994; Wauters et al., 1990; Wauters and Dhondt, 1992; Wauters et al., 1994a; Zenner et al., 2014). We also nerveless information on whether species always lived in social groups ('obligately social') or whether species were only social during specific life stages ('periodically social'). All the same, it was not possible to analyse these data as experimental manipulations of pathogens, a key factor influencing the relationship between relatedness and mortality and pathogen abundances, were simply performed for ane periodically social species (Rana latastei).
Information limitations
Request a detailed protocolOur dataset highlighted that there are several key variables where data are limited and where further empirical work would be extremely useful. In item, information on the following is currently express: (1) species that typically live with non-kin (r: kin = 41, non-kin = xv. LnCVR: kin = eighteen, non-kin = 7); (2) studies that quantify the outcome of relatedness on rates of mortality in the absenteeism of pathogens, peculiarly under natural conditions. Out of 75 studies, pathogens were excluded in xvi laboratory studies and no studies tried to explicitly exclude pathogens under field conditions. For LnCVR, pathogens were only excluded in seven laboratory studies out of a total of 32 studies; and (three) variation across groups in rates of bloodshed and pathogen abundance (out of 210 mean effect sizes, variance could only be examined in 106).
Statistical analysis
General techniques
Request a detailed protocolData were analysed using Bayesian Phylogenetic Multi-level Meta-regressions (BPMM) with Markov chain Monte Carlo (MCMC) interpretation and Gaussian error distributions in R bundle MCMCglmm (Hadfield, 2010). Data points were weighted by the inverse sampling variance associated with each of the effect size using the 'mev' term in MCMCglmm.
where northward corresponds to the number of groups, L and H are groups with low and high relatedness, respectively. Unfortunately, the difference in relatedness betwixt low and high relatedness treatments could not be included equally a moderator in analyses considering exact estimates of relatedness were non ever given (eastward.g. monogamous versus polyandrous breeders) or comparable beyond studies (east.m. estimates of relatedness from molecular markers practice non e'er equate to relatedness estimates from pedigrees/breeding designs).
The non-independence of data arising from multiple effect sizes per study were modelled by including study as a random effect. In one written report (Reber et al., 2008), there were 3 relatedness treatment groups (low, intermediate, and high) assuasive effect sizes betwixt low and intermediate, and high and intermediate to be calculated. Notwithstanding, we excluded comparisons with the intermediate treatment to avoid non-independence of issue sizes within studies (Noble et al., 2017). The non-independence of data arising from shared ancestry were modelled by including a phylogenetic variance-covariance matrix of species relationships every bit a random outcome. The phylogenetic variance-covariance matrix was created from hierarchical taxonomic classifications using the 'as.phylo' function in the R package 'ape' (see Figure 1). We also created a phylogeny using information from the open up tree of life (Rees and Cranston, 2017) using the R package 'rotl' (Michonneau et al., 2016). This produced a tree that was extremely similar, but several mollusc species were missing and we therefore used the tree created from taxonomy. Branch lengths were estimated using Grafen's method (Grafen, 1989) implemented in the R package 'ape' (Paradis, 2012).
Stock-still furnishings were considered pregnant when 95% credible intervals did not overlap with 0 and pMCMC were less than 0.05 (pMCMC = per centum of iterations above or below a test value correcting for the finite sample size of posterior samples). Default stock-still effect priors were used (independent normal priors with goose egg mean and large variance [1010]) and for random effects inverse gamma priors were used (V = 1, nu = 0.002). Each analysis was run for 1,100,000 iterations with a burn down-in of 100,000 and a thinning level of g. Convergence was checked by running each model 3 times and examining the overlap of traces, levels of autocorrelation, and testing with Gelman and Rubin's convergence diagnostic (potential scale reduction factors <1.1).
Specific analyses
Request a detailed protocolTwo sets of analyses were conducted, one on the effect of relatedness on mean rates of bloodshed and pathogen abundances (Zr) and ane on variances (LnCVR). All models were fitted with a Gaussian mistake distribution, study, species, and phylogeny as random effects and each data point was weighted by the inverse sampling variance. Half dozen analyses of mean effect sizes were conducted that had the following fixed furnishings (moderators): (1) intercept-but model to test whether overall relatedness increased susceptibility to pathogens and increased bloodshed; (2) three-level factor of whether mortality was measured in the presence of pathogens, mortality was measured in the absenteeism of pathogens, or whether the abundance of pathogens was examined (referred to here equally 'fettle measure'); (iii) four-level factor of the upshot of presence and absence of pathogens in experimental versus observational studies; (4) four-level gene of the upshot of experimentally manipulating or observing relatedness in the presence and absenteeism of pathogens; and (five) viii-level factor of the effect of living with kin and non-kin in the presence and absence of pathogens in experimental and observational studies. All analyses were repeated for LnCVR apart from 5, as variance estimates were only available for seven species that live with non-kin.
Verification analyses
Request a detailed protocolWe checked the robustness of our results to potential inbreeding furnishings (Zr and LnCVR: Supplementary file i—Tables S14 and S15), whether studies were conducted in laboratories or under natural atmospheric condition (Zr and LnCVR: Supplementary file 1—Tables S16 and S17), and the blazon of statistical tests used in studies (Zr: Supplementary file i—Table S18). To cheque for effects of potential inbreeding, nosotros repeated assay iv (see above) removing data points where at that place was any chance of inbreeding (run across Supplementary file ane—Tabular array S1 for consequence size details. Come across Supplementary file 1—Tables S14-16 for re-analysis). At that place was a large overlap in whether studies were conducted in laboratories and whether they were observational or experimental: All studies conducted in laboratories were experimental whereas for observational studies 141 outcome sizes were from field studies and 23 from laboratory studies. To bank check for laboratory furnishings, nosotros therefore restricted data to observational studies and tested if outcome sizes differed between laboratory and field studies (Supplementary file 1—Tables S16 and S17). To examine the influence of the type of statistical tests used in studies (number of different assay techniques = xv), we included 'analysis technique' as a random effect in our main model (analysis iv above: see R script 'ZrModels' M9). The main conclusions of our study remained unchanged and quantitatively similar in all verification analyses (Supplementary file ane—Tables S14-S18).
Testing for publication bias
Request a detailed protocolPublication bias across studies was checked using funnel plot visualisation and Egger'south regression (Egger et al., 1997). Egger's regressions of both Zr and LnCVR were performed by including the inverse sampling variance as a covariate in our full model (analysis 4 above: meet R script 'PublicationBias'). In both analyses, the slope of the changed sampling variance was not significantly different from zip (BPMM: inverse sampling variance on Zr CI = −0.03 to 0.01 and LnCVR CI = −0.04 to 0.12) and funnel plots of residuals were besides generally symmetrical (Figure 1—figure supplement 3; Figure 4—figure supplement 2), indicating there was little evidence of publication bias.
Data availability
All data, lawmaking and supplementary information are available at the open scientific discipline framework (OSF): http://doi.org/10.17605/OSF.IO/Q3ANE.
References
-
Book
Pair-bondingmating systems and hormones
In: Choe C. J, editors. Encyclopedia of Creature Behavior. Elsevier. pp. 611–617.
- Google Scholar
-
Briefing
Breeding habits of Twenty-Two species of marine mollusca
Proceedings of the Royal Social club New Zealand. pp. 152–174.
- Google Scholar
-
Perspectives on the beliefs of entomopathogenic Nematodes from dispersal to reproduction: traits contributing to nematode fitness and biocontrol efficacy
Journal of Nematology 44:177–184.
- PubMed
- Google Scholar
-
Book
Kinship, Recognition, Illness, and Intelligence: Constraints of Social Evolution in Fauna Societies: Theories and Facts
Nihon Scientific Guild Press.
- Google Scholar
-
Book
Meta-Assay: Accumulating Results Across Research Domains
The University of Direction Review.
- Google Scholar
Determination alphabetic character
Acceptance summary:
Group living may be beneficial for many reasons, but has costs in terms of increased rates of parasitism, in detail if grouping members are highly related. In this meta-analysis, many original studies on questions related to parasitism, relatedness and group living are brought together in 1 unifying framework. The authors conclude that living in groups tin can indeed facilitate the spread of infectious diseases, but that these costs tin be outweighed past the benefits of group living.
Determination letter after peer review:
Thank you lot for submitting your article "Living with relatives offsets the harm caused by pathogens in natural populations" for consideration by eLife. Your article has been reviewed by ii peer reviewers, one of whom is a member of our Board of Reviewing Editors, and the evaluation has been overseen by Christian Rutz as the Senior Editor. The following individual involved in the review of your submission has agreed to reveal their identity: Jacobus J Boomsma (Reviewer #2).
The reviewers take discussed their reviews with one another, and the Reviewing Editor has drafted this conclusion alphabetic character to help you prepare a revised submission.
This meta-assay addresses a double-edged sword in evolutionary biology. Group living may be beneficial for many reasons, just has costs in terms of increased rates of parasitism. Furthermore, if groups are highly related, parasites that are genetically able to infect one member of the group, may exist able to infect all of them, putting the unabridged grouping at risk. In the nowadays meta-analysis, many original studies on questions related to parasitism, relatedness and group living are brought together in one unifying framework. The authors find that, indeed, group living tin facilitate the spread of infectious diseases. However, they likewise discover that the negative effects of disease can be overcompensated by the benefits of being social. The authors stress that experimental studies are necessary to disentangle these effects. The study is of a high standard and well-conducted. The take dwelling house bulletin is articulate and of general interest.
The referees have a number of useful suggestions to aid y'all improve the manuscript. From our perspective, all of them seem easy to incorporate and there are no major bug. A detailed list is given below.
Reviewer #1 (Recommendations for the authors):
I was surprised to read that among the primal words used at different stages of the refinement procedure for the literature search, term like: clone, clonal, social,.… did not show up.
A focus of the study is on effects on mortality. However, many pathogens are hardly lethal, just reduce other fitness relevant traits strongly, such as competitive power, work performance, sexual attractiveness and fecundity.
Hand-picked examples of private studies read squeamish and are entertaining, but they do not support the overall determination, but rather bias the observer. I suggest to leave them out, unless of specific relevance. E.g. the example of the frog and firm mouse at the beginning of the result section, is not needed. Likewise the Tribolium and the worm examples (line 149).
Reviewer #2 (Recommendations for the authors):
– 38. I have non checked but remember that Baer and Schmid-Hempel (you lot cite these papers) had a more direct experimental manipulation approach than Liersch and Schmid-Hempel considering they inseminated bumblebee queens with either one or 4 ejaculates. Worth citing here likewise?
– 44. Why is extinction chance relevant? Natural selection simply sees private inclusive fettle. Rephrase to avoid ambiguity?
– 57-60. Unclear sentence – rephrase.
– 67. Information technology was Sylvia Cremer who developed the field of 'social immunity' in the first decade of this century. I think that pioneering work should be cited here. You have 1 of her papers (Ugelvig et al), but peradventure the review past Cremer and Sixt is more than full general.
– 79. Replace the second 'to' by 'as in' to make judgement clearer?
– 106. It seems rather excessive to use three decimals here.
– 191-193. As in 44, the phrasing seems to take a slight and I'm sure unintentional group-option camber. Tin some rewording repair that?
– 205-207In Acromyrmex individuals are non-totipotent caste members and they exercise non have a 'choice' between living with relatives of not-relatives. Farther, relatedness in Acromyrmex colonies is very low for ant standards, then even though you have them in the loftier relatedness (>0.25) category, they would exist in the low-relatedness category when you would focus on social Hymenoptera only. The Hughes papers that you cite accept precise relatedness estimates I believe. Somehow the phrasing of your text does not seem to capture these peculiarities.
– The numerical order of the supplementary tables seems rather haphazard. Should they not be numbered in the order they are presented in the text?
https://doi.org/10.7554/eLife.66649.sa1
Writer response
Reviewer #one (Recommendations for the authors):
I was surprised to read that among the key words used at different stages of the refinement procedure for the literature search, term like: clone, clonal, social,.… did not show upwardly.
We initially tried combinations of search terms to identify those that increased the number of relevant papers without inflating the number of search hits. Relevant papers that used clone and clonal besides used genetic variety and monoculture and were therefore captured by our searches, but returned many hits from agricultural studies which we wanted to exclude. The term social was too broad, dramatically increasing our search hits. Relevant papers that used social also used the terms group, colony or relatedness then nosotros are confident that are search criteria captured relevant papers associated with these extra key words.
We have now clarified this point in the methods, section 'Literature searches' which reads:
"Initial exploration of search terms included other words ('clone','clonal','social'). Even so, these terms inflated the number of search hits and papers with relevant information were retrieved using other terms included in our search criteria ('group', 'colony' or 'relatedness')" (Lines 276-279).
A focus of the study is on effects on mortality. Nevertheless, many pathogens are hardly lethal, but reduce other fitness relevant traits strongly, such equally competitive power, work performance, sexual attractiveness and fecundity.
We agree that pathogens can accept varied effects on unlike fettle related traits across bloodshed that are probable important for individuals. Notwithstanding, the relationship betwixt within-group relatedness and measures such as fecundity tin vary for many reasons other than pathogens making interpretation complicated. For example, local mate contest theory predicts that investment in sexual traits varies with local relatedness amid individuals. Furthermore, traits such as sexual attractiveness and competitive ability are not hands interpreted in some species. In social insects, the relationship between relatedness and average attractiveness or competitive ability of workers does not take clear meaning to the hypotheses we tested. Nosotros therefore decided to stick with measures (bloodshed and pathogen abundance) that clearly related to the hypotheses we tested and that were comparable across all species.
We have at present antiseptic in the methods why we excluded studies using other proxies of fitness. It reads:
"We did not include studies examining the relationship betwixt within-grouping relatedness and other fitness related measures, such every bit fecundity or competitive ability, because such measures are influenced by factors other than pathogens" (Lines293-295).
Hand-picked examples of individual studies read nice and are entertaining, simply they do not support the overall decision, but rather bias the observer. I suggest to leave them out, unless of specific relevance. Eastward.g. the example of the frog and house mouse at the starting time of the result department, is non needed. Besides the Tribolium and the worm examples (line 149).
Nosotros are grateful for the proposition. Nosotros did not intend to bias the observer, just to give the reader greater insight into the primary studies. Meta-analyses tin can help cut through the species-specific idiosyncrasies allowing more than general interpretation, simply ane downside is that the reader is left wondering what kind of studies were included. For this reason, nosotros believe some examples tin can increase interpretability. Yet, nosotros have reduced these (example at the start of the results removed) in accordance with the referee'due south suggestion.
Reviewer #two (Recommendations for the authors):
– 38. I take non checked but remember that Baer and Schmid-Hempel (you lot cite these papers) had a more than direct experimental manipulation approach than Liersch and Schmid-Hempel because they inseminated bumblebee queens with either 1 or 4 ejaculates. Worth citing here too?
We have now added this citation.
– 44. Why is extinction take a chance relevant? Natural choice only sees individual inclusive fitness. Rephrase to avert ambiguity?
We take now rephrased this, which reads:
"What remains unclear, however, is whether this translates into college rates of bloodshed, or whether the benefits of living with relatives are large enough to first the costs of increased illness risk" (Lines 49-51).
– 57-sixty. Unclear judgement – rephrase.
We have revised this sentence, which reads:
"For instance, a negative relationship betwixt relatedness and the abundance of pathogens tin occur either because groups of relatives are less susceptible to pathogens, or conversely because groups of relatives frequently die from pathogens and so rarely observed" (Lines 62-65).
– 67. It was Sylvia Cremer who adult the field of 'social immunity' in the start decade of this century. I think that pioneering work should exist cited here. You lot have one of her papers (Ugelvig et al), but possibly the review past Cremer and Sixt is more general.
Give thanks you lot for the suggestion. Nosotros at present cite Ugelvig et al. as well as Cremer and Sixt.
– 79. Replace the second 'to' by 'every bit in' to brand sentence clearer?
Addressed.
– 106. It seems rather excessive to utilise 3 decimals here.
Agreed! The manuscript is written in R markdown and then results can be directly inserted into text to increase reproducibility, but sometimes the formatting slips through the net. Nosotros accept now inverse the results to 2 decimals throughout.
– 191-193. Equally in 44, the phrasing seems to have a slight and I'm sure unintentional grouping-selection slant. Tin some rewording repair that?
We have now rephrased this sentence which reads:
"I explanation is that individuals answer to greater pathogen pressure by forming more than genetically diverse groups" (Lines 210-211).
– 205-207: In Acromyrmex individuals are non-totipotent degree members and they practise not have a 'choice' betwixt living with relatives of non-relatives. Further, relatedness in Acromyrmex colonies is very low for ant standards, and then even though you accept them in the high relatedness (>0.25) category, they would be in the low-relatedness category when you would focus on social Hymenoptera only. The Hughes papers that you cite take precise relatedness estimates I believe. Somehow the phrasing of your text does not seem to capture these peculiarities.
This example was to illustrate that spatial segregation and task sectionalisation within groups tin reduce the spread of pathogen manual among group members. While Acromyrmex have lower (derived) relatedness than some other social insects, they still have relatively high relatedness compared to the other species in our dataset (E.g. Acromyrmex octospinosus: r = ca. 0.33, Boomsma et al. 1999. Acromyrmex echinatior r = 0.380±0.042 Sumner et al. 2004). Nosotros take tried to rephrase this section to clarify the point of this example.
Information technology reads:
"For example, in leaf cutter ants, Acromyrmex spp, workers exterior the colony, where pathogens are more than prevalent, do not enter the inner colony (Camargo et al., 2007). Contamination of food past pathogens is also limited by workers outside the colony performing dedicated tasks, such equally foraging versus waste management (Waddington and Hughes, 2010)" (Lines 200-204).
– The numerical gild of the supplementary tables seems rather haphazard. Should they not be numbered in the lodge they are presented in the text?
Thank yous for pointing this out. The supplementary tables are at present numbered in the guild they announced in the text.
https://doi.org/10.7554/eLife.66649.sa2
Article and author information
Author details
Funding
Knut och Alice Wallenbergs Stiftelse (2018.0138)
- Charlie Kinahan Cornwallis
Vetenskapsrådet (2017-03880)
- Charlie Kinahan Cornwallis
The funders had no office in report blueprint, information collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
This research was funded by the Knut and Alice Wallenberg Foundation (Wallenberg Academy fellowship number 2018.0138) and the Swedish Research Council (grant number 2017–03880). We are very grateful to Dan Noble, Dieter Ebert, Christian Rutz, Jacobus Boomsma, and an anonymous reviewer for comments on the manuscript.
Senior Editor
- Christian Rutz, Academy of St Andrews, Uk
Reviewing Editor
- Dieter Ebert, University of Basel, Switzerland
Reviewer
- Jacobus J Boomsma, University of Copenhagen, Denmark
Publication history
- Received: January 18, 2021
- Accepted: June 19, 2021
- Version of Tape published: July 26, 2022 (version 1)
Copyright
© 2021, Bensch et al.
This article is distributed under the terms of the Artistic Commons Attribution License, which permits unrestricted use and redistribution provided that the original writer and source are credited.
Metrics
-
- 428
- Page views
-
- 45
- Downloads
-
- 1
- Citations
Commodity commendation count generated by polling the highest count across the following sources: Crossref, PubMed Central, Scopus.
Download links
A ii-function list of links to download the article, or parts of the commodity, in diverse formats.
Downloads (link to download the article as PDF)
Download citations (links to download the citations from this article in formats compatible with various reference manager tools)
Open up citations (links to open up the citations from this article in various online reference director services)
Source: https://elifesciences.org/articles/66649

0 Response to "Why Can Evolution in Mosquitoes Be More Easily Observed Than Evolution in a Population of Squirrels?"
Post a Comment